You Don’t Need to Learn All the Weights on tabular data: The Case for rvflnet (a nonlinear expressive glmnet) on regression, classification and survival analysis
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.
Introduction
Random Vector Functional Link (RVFL) networks offer a simple yet powerful alternative to traditional neural networks for tabular data. Instead of learning hidden layers through backpropagation, RVFL generates them randomly (or not, if using a deterministic sequence of quasi-random numbers) and focuses all learning effort on a final, regularized linear model.
Formally, let
\[X \in \mathbb{R}^{n \times p}\]be the input data. RVFL networks (the ones described in this blog post) construct a set of nonlinear features by projecting (X) onto a random matrix
\[W \in \mathbb{R}^{p \times m},\]and applying an activation function (\(g(\cdot)\)):
\[H = g\left( \frac{X – \mu}{\sigma} ; W \right).\]These random nonlinear features are then concatenated with the original inputs to form an augmented design matrix:
\[Z = [X | H].\]The model prediction is obtained by fitting a linear model on this expanded space (hence, a nonlinear GLM):
\[\hat{y} = Z \beta.\]Because (Z) can be high-dimensional and highly redundant, RVFL networks (the ones described in this blog post) rely on Elastic Net regularization (glmnet) to estimate the coefficients:
In this framework, randomness creates a rich pool of nonlinear transformations, while regularization selects and stabilizes the most useful ones. The result is a nonlinear model that combines the flexibility of neural networks with the efficiency and robustness of linear methods.
Of course, this blog post is not a proof of the title. It’s about R package rvflnet. But you can appreciate the high performance of RVFLs on regression, classification and survival analysis, an notably on the controversial Boston dataset (performs on par with Random Forest or Gradient Boosting).
0 – Install package
install.packages("survival", repos = "https://cran.r-project.org") # survival analysis
install.packages("remotes", repos = "https://cran.r-project.org")
devtools::install_github('thierrymoudiki/rvflnet') # Nonlinear glm (RVFL networks)
1 – Regression
set.seed(123)
library(glmnet)
data(Boston, package = "MASS")
# -------------------------
# Data
# -------------------------
X <- as.matrix(Boston[, -14])
y <- Boston$medv
n <- nrow(X)
idx <- sample(1:n, size = round(0.8 * n))
X_train <- X[idx, ]
y_train <- y[idx]
X_test <- X[-idx, ]
y_test <- y[-idx]
# -------------------------
# Grid
# -------------------------
grid <- expand.grid(
n_hidden = c(175, 200, 225, 250),
alpha = seq(0.1, 0.5, by=0.2),
include_original = c(TRUE, FALSE),
seed = 1,
stringsAsFactors = FALSE
)
results <- vector("list", nrow(grid))
# -------------------------
# Loop
# -------------------------
for (i in seq_len(nrow(grid))) {
params <- grid[i, ]
#cat("\n========================================\n")
#cat(sprintf("Run %d / %d\n", i, nrow(grid)))
#print(params)
# -------------------------
# Fit model
# -------------------------
fit <- rvflnet::rvflnet(
X_train, y_train,
n_hidden = params$n_hidden,
activation = "sigmoid",
W_type = "gaussian",
seed = params$seed,
include_original = params$include_original, # direct link, skip connection or not
alpha = params$alpha
)
# -------------------------
# Evaluate full lambda path
# -------------------------
lambdas <- fit$fit$lambda
preds <- predict(fit, newx = X_test, s = lambdas)
rmse_path <- sqrt(colMeans((preds - y_test)^2))
best_idx <- which.min(rmse_path)
best_rmse <- rmse_path[best_idx]
best_lambda <- lambdas[best_idx]
# -------------------------
# Sparsity
# -------------------------
coef_mat <- coef(fit, s = best_lambda)
nonzero <- sum(coef_mat[-1, 1] != 0)
# -------------------------
# Verbose output
# -------------------------
#cat(sprintf("Best RMSE: %.4f\n", best_rmse))
#cat(sprintf("Best lambda: %.6f\n", best_lambda))
#cat(sprintf("Non-zero coeffs: %d\n", nonzero))
# -------------------------
# Store
# -------------------------
results[[i]] <- data.frame(
n_hidden = params$n_hidden,
alpha = params$alpha,
include_original = params$include_original,
seed = params$seed,
rmse = best_rmse,
lambda = best_lambda,
nonzero = nonzero
)
}
# -------------------------
# Aggregate
# -------------------------
results_df <- do.call(rbind, results)
results_df <- results_df[order(results_df$rmse), ]
print(head(results_df))
Loading required package: Matrix
Loaded glmnet 4.1-10
n_hidden alpha include_original seed rmse lambda nonzero
s= 0.027561759 200 0.1 TRUE 1 2.881935 0.02756176 190
s= 0.017620327 200 0.3 TRUE 1 2.884739 0.01762033 167
s= 0.012734248 200 0.5 TRUE 1 2.889339 0.01273425 158
s= 0.036435024 175 0.1 TRUE 1 2.920012 0.03643502 165
s= 0.016833926 175 0.5 TRUE 1 2.938472 0.01683393 136
s= 0.023293035 175 0.3 TRUE 1 2.941267 0.02329304 144
An RMSE of 2.88 is on par with Random Forest or Gradient Boosting, with a significantly faster computation time.
2 - Classification
2 - 1 Binary Classification
set.seed(123)
data(iris)
# Binary classification: setosa vs others
y <- ifelse(iris$Species == "setosa", 1, 0)
X <- as.matrix(iris[, 1:4])
# Train/test split
n <- nrow(X)
idx <- sample(1:n, size = round(0.8 * n))
X_train <- X[idx, ]
y_train <- y[idx]
X_test <- X[-idx, ]
y_test <- y[-idx]
# -------------------------
# Fit model
# -------------------------
cv_model <- rvflnet::cv.rvflnet(
X_train, y_train,
n_hidden = 50,
activation = "relu",
W_type = "gaussian",
family = "binomial",
nfolds = 5
)
# -------------------------
# Predictions (probabilities)
# -------------------------
(probs <- predict(cv_model, X_test, type = "response"))
# Convert to class
y_pred <- ifelse(probs > 0.5, 1, 0)
all.equal(as.numeric(y_pred), as.numeric(predict(cv_model, X_test, type="class")))
# -------------------------
# Diagnostics
# -------------------------
# Accuracy
acc <- mean(drop(y_pred) == y_test)
cat("Accuracy:", acc, "\n")
# Confusion matrix
table(Predicted = y_pred, Actual = y_test)
| lambda.min |
|---|
| 0.9997617002 |
| 0.9992267955 |
| 0.9997120678 |
| 0.9997524867 |
| 0.9996600481 |
| 0.9992472082 |
| 0.9996101744 |
| 0.9999356520 |
| 0.9998139568 |
| 0.9995418762 |
| 0.0003328885 |
| 0.0003328885 |
| 0.0003328885 |
| 0.0019937012 |
| 0.0003328885 |
| 0.0005459970 |
| 0.0003328885 |
| 0.0005035848 |
| 0.0003328885 |
| 0.0003328885 |
| 0.0003328885 |
| 0.0003328885 |
| 0.0003328885 |
| 0.0003328885 |
| 0.0003328885 |
| 0.0003328885 |
| 0.0003328885 |
| 0.0003328885 |
| 0.0003328885 |
| 0.0003328885 |
TRUE
Accuracy: 1 Actual Predicted 0 1 0 20 0 1 0 10
2 - 2 Multiclass Classification
set.seed(123)
data(iris)
y <- as.numeric(iris$Species)
X <- as.matrix(iris[, 1:4])
# Train/test split
n <- nrow(X)
idx <- sample(1:n, size = round(0.8 * n))
X_train <- X[idx, ]
y_train <- y[idx]
X_test <- X[-idx, ]
y_test <- y[-idx]
# -------------------------
# Fit model
# -------------------------
cv_model <- rvflnet::rvflnet(
X_train, y_train,
n_hidden = 50,
activation = "relu",
W_type = "gaussian",
family = "multinomial",
nlambda = 25,
nfolds = 5
)
# -------------------------
# Diagnostics
# -------------------------
# Accuracy
acc <- colMeans(predict(cv_model, X_test, type="class") == y_test)
cat("Accuracies:", acc, "\n") # consider other metrics
Accuracies: 0.1666667 0.7666667 0.9333333 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667 0.9666667
3 - Nonlinear Cox survival analysis
3 - 1 Example 1
library(survival)
library(rvflnet)
data(ovarian)
X <- as.matrix(ovarian[, c("age", "resid.ds", "rx", "ecog.ps")])
y <- Surv(ovarian$futime, ovarian$fustat)
set.seed(123)
n <- nrow(X)
train_idx <- sample(1:n, size = round(0.8 * n))
X_train <- X[train_idx, ]
X_test <- X[-train_idx, ]
y_train <- y[train_idx]
y_test <- y[-train_idx]
# -------------------------
# Fit model
# -------------------------
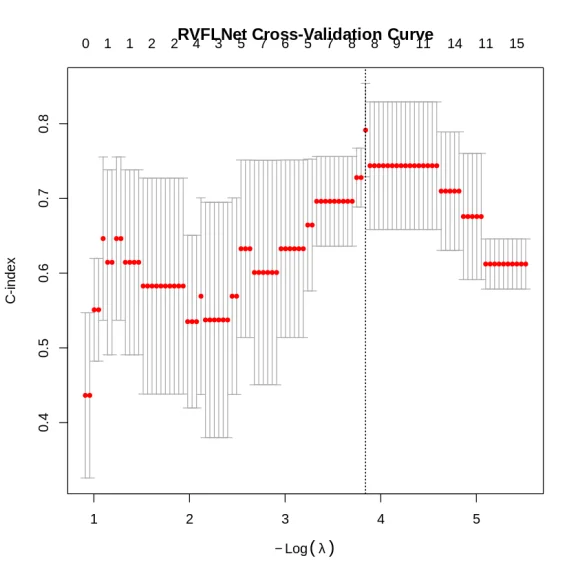
cv_fit <- rvflnet::cv.rvflnet(
X_train, y_train,
family = "cox",
nfolds = 5,
type.measure = "C"
)
plot(cv_fit)
# Out-of-sample C-index
print(glmnet::Cindex(pred = predict(cv_fit, X_test), y = y_test))
Warning message in data(ovarian):
“data set ‘ovarian’ not found”
[1] 0.8571429
3 - 2 Example 2
library(glmnet)
library(survival)
data(pbc)
pbc2 <- pbc[!is.na(pbc$trt), ]
pbc2$event <- as.integer(pbc$status[!is.na(pbc$trt)] == 2)
pbc2$sex_n <- as.integer(pbc2$sex == "f")
feat_cols <- c("trt","age","sex_n","ascites","hepato","spiders","edema",
"bili","chol","albumin","copper","alk.phos","ast",
"trig","platelet","protime","stage")
df <- pbc2[, c("time", "event", feat_cols)]
for (col in feat_cols)
if (any(is.na(df[[col]])))
df[[col]][is.na(df[[col]])] <- median(df[[col]], na.rm = TRUE)
set.seed(42)
idx_train <- sample(nrow(df), floor(0.75 * nrow(df)))
train <- df[idx_train, ]; test <- df[-idx_train, ]
X_tr <- as.matrix(train[, feat_cols])
X_te <- as.matrix(test[, feat_cols])
y_tr <- Surv(train$time, train$event)
fit <- rvflnet::rvflnet(
X_tr, y_tr,
family = "cox",
alpha=0.1, lambda=0.1 # not recommended
)
y_te <- Surv(test$time, test$event)
ci <- glmnet::Cindex(predict(fit, X_te), y_te)
cat("\n=== Test-set C-index ===\n")
print(ci)
=== Test-set C-index ===
[1] 0.8218117
fit <- rvflnet::rvflnet(
X_tr, y_tr,
family = "cox",
alpha=0.1, nlambda=50
)
y_te <- Surv(test$time, test$event)
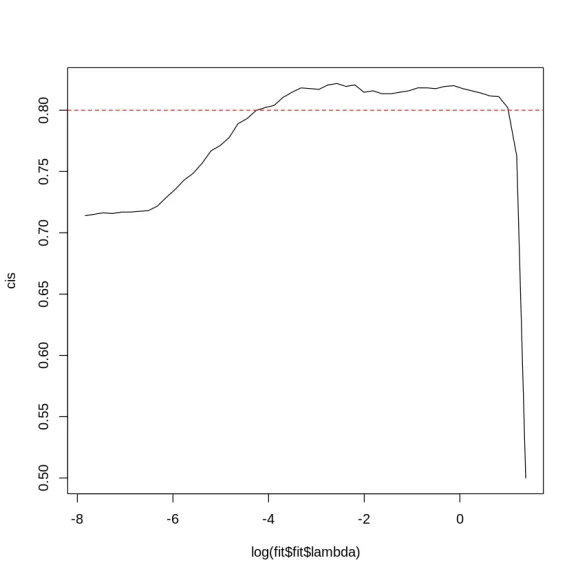
(cis <- apply(predict(fit, X_te), 2, function(x) glmnet::Cindex(x, y_te)))
#cat("\n=== Test-set C-index ===\n")
plot(log(fit$fit$lambda), cis, type = 'l')
abline(h=0.8, lty=2, col="red")
- s0
- 0.5
- s1
- 0.762812872467223
- s2
- 0.802145411203814
- s3
- 0.811084624553039
- s4
- 0.811680572109654
- s5
- 0.814064362336114
- s6
- 0.815852205005959
- s7
- 0.817640047675805
- s8
- 0.820023837902265
- s9
- 0.81942789034565
- s10
- 0.817640047675805
- s11
- 0.81823599523242
- s12
- 0.81823599523242
- s13
- 0.815852205005959
- s14
- 0.814660309892729
- s15
- 0.813468414779499
- s16
- 0.813468414779499
- s17
- 0.815852205005959
- s18
- 0.814660309892729
- s19
- 0.82061978545888
- s20
- 0.81942789034565
- s21
- 0.82181168057211
- s22
- 0.82061978545888
- s23
- 0.817044100119189
- s24
- 0.817640047675805
- s25
- 0.81823599523242
- s26
- 0.814660309892729
- s27
- 0.810488676996424
- s28
- 0.803933253873659
- s29
- 0.802145411203814
- s30
- 0.799761620977354
- s31
- 0.793206197854589
- s32
- 0.789034564958284
- s33
- 0.777711561382598
- s34
- 0.771156138259833
- s35
- 0.766984505363528
- s36
- 0.756853396901073
- s37
- 0.748510131108462
- s38
- 0.743146603098927
- s39
- 0.735399284862932
- s40
- 0.728843861740167
- s41
- 0.721692491060787
- s42
- 0.718116805721096
- s43
- 0.717520858164482
- s44
- 0.716924910607867
- s45
- 0.716924910607867
- s46
- 0.715733015494636
- s47
- 0.716328963051251
- s48
- 0.715137067938021
- s49
- 0.713945172824791
R-bloggers.com offers daily e-mail updates about R news and tutorials about learning R and many other topics. Click here if you're looking to post or find an R/data-science job.
Want to share your content on R-bloggers? click here if you have a blog, or here if you don't.

